This is our blog series interviewing our 2019 Google Summer of Code students, who will be working remotely for InterMine for 3 months on a variety of projects. We’ve interviewed Akshat Bhargava, who will be creating data visualisations for BlueGenes.
Hi Akshat! We’re really excited to have you on board as part of the team this summer. Can you introduce yourself?
 Hi InterMine team, I’m very excited too for this upcoming summer! I’m a Computer Science undergraduate going to start my 3rd year this August. I’m primarily a Javascript Developer (Web & Hybrid Mobile) and have been working with it for the last 2.5+ years, but the real me is a person who loves to solve problems in general, may they be related to programming or not. I’ve been exploring the field of data visualization for the last few months and I am in love with it. Have a look at IPL (cricket) data viz I created a few months back here.
Hi InterMine team, I’m very excited too for this upcoming summer! I’m a Computer Science undergraduate going to start my 3rd year this August. I’m primarily a Javascript Developer (Web & Hybrid Mobile) and have been working with it for the last 2.5+ years, but the real me is a person who loves to solve problems in general, may they be related to programming or not. I’ve been exploring the field of data visualization for the last few months and I am in love with it. Have a look at IPL (cricket) data viz I created a few months back here.
Apart from coding, I love reading about psychology, history and watching horror movies.
What interested you about GSoC with InterMine?
I feel it magical how numbers show their true faces when seen via a meaningful visualization, and this is why I’m most excited for this summer with InterMine.
Real World Bio Data + Data Viz = Something big coming in! ❤
Another reason for my interest in InterMine, is that I applied to InterMine last year too for Cross InterMine Search Tool and couldn’t make it, but understood it’s community and how they work. The mentors are very helpful and supportive to everyone, so I directly jumped here this year. 😀
Tell us about the project you’re planning to do for InterMine this summer.
InterMine has tons of different types of biological data, this summer I’ll mostly be working on discussing and developing visualizations for data, making it easier to biologists to understand it in a easier way, and draw relevant conclusions with a single sight to the graphs.
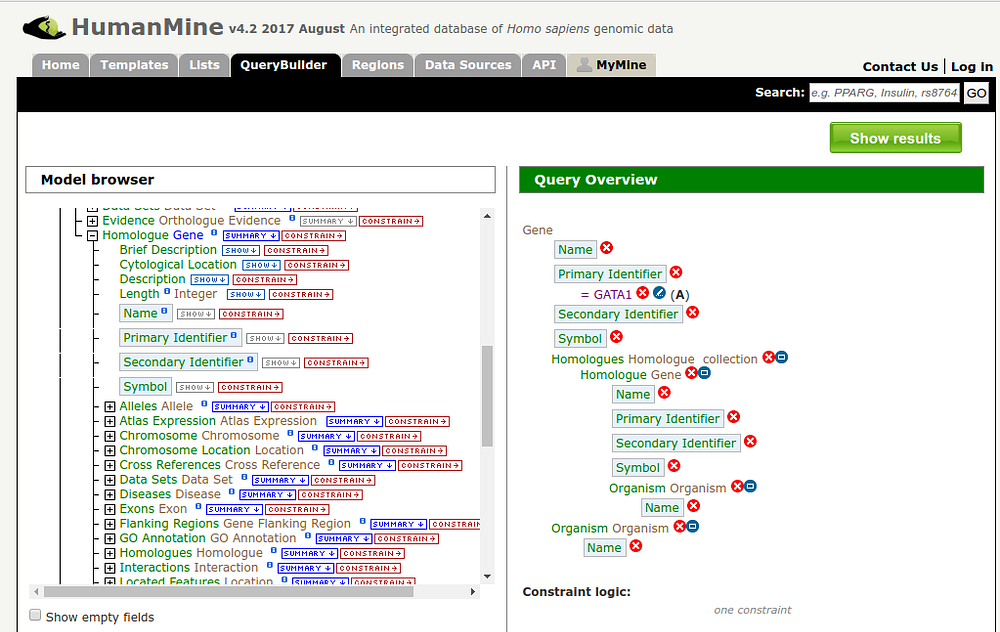
There is a software called BlueGenes, which is already developed and helps explore different mines, it provides a tool API which allows Javascript developers to create additional visualization tools on top of it, which can be integrated on any Gene or Protein result page. My goal for this summer is to create a different variety of such visualization tools in order to enrich the visualization of different types of data.
As an example of how useful is what I’m doing, one of the visualizations I’ll be developing will help us understand how the expression of a particular gene is distributed among different tissues. This information is helpful for cancer biologists that want to assess if a gene is highly expressed across different tissues of an organism, because that gives a relative picture on to what degree it’s implicated in diseases.
Are there any challenges you anticipate for your project? How do you plan to overcome them?
Data visualization is something that requires you to understand the data properly first in order to be able to actually create some meaningful visualization out of it, and since I’m not very familiar with InterMine’s data model and the related biological terms, I’ll face some difficulties during my thought process of “what and why” to visualize. To overcome this, I’m already exploring more and more of the InterMine’s data model, trying to understand how to deal with different types of data, and how to create the appropriate visualization for them. Mentors are really helping me out with this (overall in terms of tech, viz and everything). 🙂
Share a meme or gif that represents your project





You must be logged in to post a comment.